GSoC 2025: Final Report
This summer has been one of the most eventful and rewarding experiences so far, filled with incredible learning. I had the chance to dive deep into research and development, and I’m excited to continue thriving in the dynamic world of data science
In this post I will walk you through all the work I have done over this summer as a part of GSoC 2025 with the PEcAn Project.
Project Summary
Organization: PEcAn projects
Mentors: David LeBauer, Nihar Sanda, Henry Priest
Student: Akash B V
Project Title : Development of Data Preparation Workflows for Ensemble-Based Modeling
Introduction
Ecosystem model predictions are inherently uncertain due to four primary sources of variability that propagate through model simulations:
1. Initial Conditions : Uncertainty in starting ecosystem states (soil carbon, vegetation biomass, nutrient pools)
2. Weather Data : Measurement errors and spatial interpolation uncertainties in meteorological forcing
3. Soil Properties : Spatial heterogeneity and measurement limitations in soil characteristics
4. Model Parameters : Imperfect knowledge of ecosystem process rates and relationships
Traditional modeling approaches using single "best estimate" inputs fail to capture this uncertainty, leading to overconfident predictions that cannot support robust decision-making. Ensemble modeling approaches address this limitation by creating multiple plausible input scenarios that represent the full range of uncertainty in each component.
This project focuses on developing modular data preparation workflows within the PEcAn ecosystem to support ecological modeling. It targets the integration of diverse data sources for meteorology (e.g, ECMWF/ERA5, AmeriFlux), soil (e.g, gSSURGO, SoilGrids, BADM), and vegetation initial conditions (e.g, LandTrendr AGB, BADM ) and generate ensemble-based initial conditions to propagate uncertainty through ecosystem model predictions. This approach is scientifically critical because ecological data inherently contains uncertainty from measurement errors, spatial heterogeneity, and temporal variability. By creating multiple plausible initial condition scenarios rather than using single-point estimates, the framework enables robust probabilistic forecasting that quantifies prediction confidence bounds and supports more informed ecological decision-making.
About PEcAn
The Predictive Ecosystem Analyzer (PEcAn) is an open-source framework designed to make ecosystem modeling more accessible, transparent, and reproducible. It serves as an ecoinformatics toolbox, providing structured workflows that integrate with various ecosystem models to streamline data synthesis, uncertainty propagation, and ecological predictions.
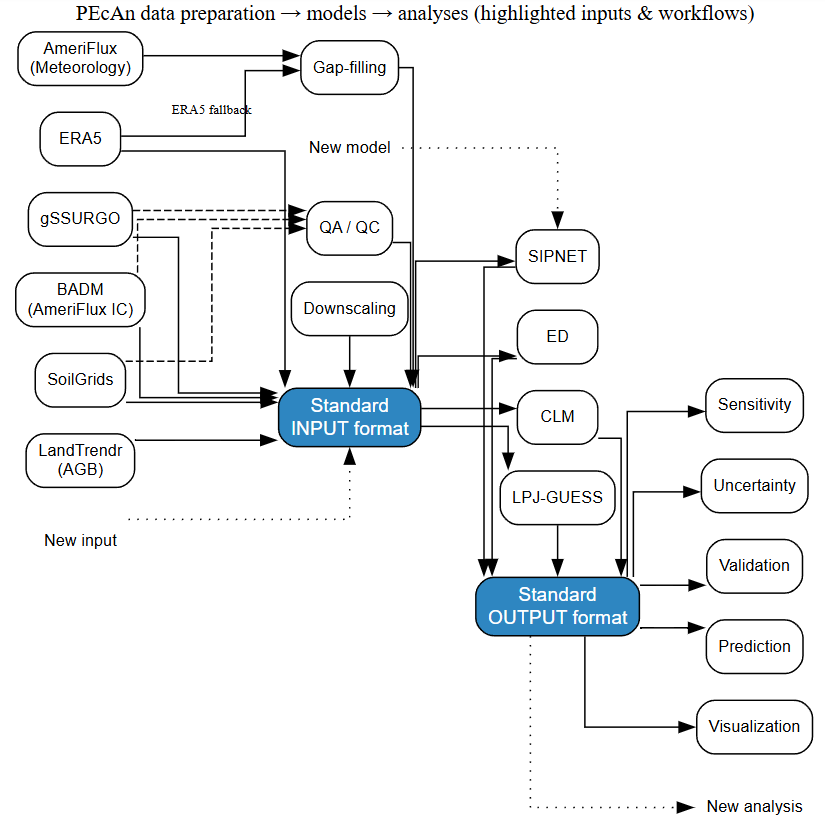
Figure 1: PEcAn data preparation workflow demonstrating community-scale efficiency through standardized input/output formats. Diverse data sources (SoilGrids, gSSURGO, AmeriFlux, ERA5, BADM, LandTrendr) are processed through unified workflows (gap-filling, downscaling, ensemble generation) into a common input format that feeds multiple ecosystem models (ED, CLM, LPJ-GUESS, SIPNET) and analyses. This approach reduces converter complexity from quadratic (n × m) to linear (n + m) scaling, enabling efficient integration of new data sources, models, and analyses without requiring pairwise conversions. Blue boxes represent the standardized formats that eliminate redundant development work across the modeling community.
Project Impact and Applications
This project directly supports critical carbon quantification efforts funded by NASA's Carbon Monitoring System and California Air Resources Board (CARB). These programs require spatially-explicit carbon inventory products with rigorous uncertainty quantification for climate policy and carbon market applications. By developing ensemble-based initial condition workflows, this work enables PEcAn to meet the explicit NASA CMS requirement that all projects include explicit plans to quantify uncertainty in any data, model, fused model-data products, or findings.
The improved data preparation workflows support:
NASA CMS carbon stock mapping : Providing uncertainty-bounded estimates of ecosystem carbon pools across landscapes
CARB compliance carbon inventories : Enabling statistically robust carbon accounting for regulatory frameworks
Climate model initialization : Supplying ensemble initial conditions that propagate data uncertainty through Earth system predictions
The workflows developed in this project including SoilGrids soil carbon ensembles, ERA5 meteorological uncertainty propagation, and BADM vegetation carbon pool processing directly provide the ensemble initial condition capabilities required by these carbon monitoring frameworks.
Pre-GSoC Period:
During this period my first priority was to review the
PEcAn.data.landandPEcAn.data.atmospherepackages, along with other essential packages required for data processing.Used this time as a buffer to finish up some of the previous opened PRs and getting merged before the start of the coding period :
Pkgdown Build Process and fix documentation (PR #3482 )
And later opened a PR for an upgrading to Bootstrap 5, plus added some nice navigation improvements and dynamic branch-specific links to source code of a package and BugReports, and some other improvements.
Pkgdown Bootstrap 5 Upgrade and Navigation Improvements (PR #3510 )
I started by understanding the different data sources that PEcAn relies on and spent time brainstorming how they integrate into the workflows.
Testing the existing functions and fixing the errors and warnings thrown by them.
Collected mentor feedback and refined workflow structures and implementation plans before the coding period began.
Work Completed during GSoC 2025
My aim in this summer to write a workflow to generate an ensembles from different data sources and to pass SOC and other IC’s to all the sources where it has been missing.
Three Types of Data + Uncertainty
1) Soil
Development of IC_SOILGRID_Utilities for SoilGrids Data Processing (PR #3508 and #3506)
I introduced a new utility script, IC_SOILGRID_Utilities.R, aimed at streamlining the processing of SoilGrids data for generating soil carbon initial condition (IC) files. This script provides key functions for extracting, processing, and generating ensemble members from SoilGrids250m data. I also fixed several reliability issues in soilgrids_soilC_extract, improving its error handling and output robustness.
Typically involves writing a new functions:
soilgrids_ic_process: Acts as a wrapper function for extracting, pre-processing, and generating ensemble members at specified soil depths (e.g , 0-30 cm, 0-200 cm), manages caching, and writes the results to NetCDF IC files.preprocess_soilgrids_data: Filters and cleans SoilGrids data, computes coefficient of variation (CV) distributions, and prepares inputs for ensemble generationgenerate_soilgrids_ensemble: Generates ensemble members for soil carbon at a given site and depth using site-specific uncertainty or integrating over CV distributions.
SOC Integration and Improvements in the gSSURGO pipeline ( PR #3534 )
I focused on integrating SOC estimation and ensemble generation in gSSURGO. I added support for spatial sampling with configurable grids (defaulting to a 3×3 grid of 100m cells), which helps capture within-site variability more realistically. Soil properties are now aggregated using area-weighted means, giving more representative results. Another key improvement was refining SOC estimation with a coarse fragment correction, ensuring that inert rock fragments don’t inflate soil carbon stock values. This is achieved using the formula ( OCS=SOC×BD×thickness×(1−coarse fraction) ). I also introduced helper functions like soc2ocs (to convert SOC concentration into carbon stock) and om2soc (to convert organic matter to SOC) that help standardize transformations across the codebase, making the workflow more flexible and reusable. Finally fixed a issues in the gSSURGO.Query function and added the chkey horizon identifier to enable coarse fragment correction in SOC estimation, along with a unit test to ensure the functionality works as intended.
2) Initial Conditions
BADM Initial Condition (IC) Processing ( PR #3536 )
I began by familiarizing myself with the AmeriFlux BADM (Biological, Ancillary, Disturbance, and Metadata) schema - site level information that describes ecosystem characteristics and carbon pools at AmeriFlux carbon monitoring stations. I reviewed the AmeriFluxer package and experimented with it to validate the data. This helped clarify the design and behavior of the BADM schema implementation.
Building on this understanding, I opened a PR to refactor and improve the BADM IC workflow, making the code more robust, schema-compliant, and maintainable. The updates addressed both technical bugs and conceptual alignment with the AmeriFlux BADM structure, ensuring correct extraction and processing of carbon pools for ensemble IC generation. The workflow now includes all relevant carbon pools (ROOT_BIOMASS, AG_BIOMASS, SOIL_STOCK, LIT_BIOMASS) . Additionally, I wrote tests to validate the extraction and processing.
3) Weather data
ERA5 is an important source of weather data from historical reanalyses as well as ensemble based forecasts
Removing Python Dependency and Improving ERA5 Data Download ( PR #3547 )
This work was necessary to adapt to major changes in ERA5 data access infrastructure, including ECMWF's new API requirements and the transition away from GRIB-to-NetCDF conversion dependencies. The updates ensure continued reliable access to ERA5 data while making the framework more robust and extensible for future API changes.
Initially, ERA5 data download relied on a Python dependency (cdsapi via reticulate)
Limitations:
Setting up an environment that uses Python from within R is cumbersome and fickle.
Data download integration and test setup required additional dependencies.
Approach used:
Removed the Python dependency and migrating to the native R ecmwfr package.
Downloaded ERA5 data directly in NetCDF format, eliminating the need for GRIB-to-NetCDF conversion.
Added support for flexible selection of product type (ensemble vs reanalysis), dataset, and time steps, instead of hardcoding values.
Developed unit tests to validate the functionality of the ERA5 download workflow.
The downloaded data was reviewed for its format and file naming to confirm both the availability of the data source and that the data aligns with PEcAn’s expected format, keeping legacy compatibility intact for downstream processing. These improvements are indeed essential and would be easiest in the long run.
Generalizing ERA5 extraction and CF conversion to support both ensemble and reanalysis data processing ( PR #3584 )
Limitations:
Previously, the
met2CF.ERA5functions assumed the presence and order of specific ERA5 variables (e.g, ssrd, tp, d2m, sp) and was fixed to a 3-hour timestep.The function attempted to calculate specific humidity even when required variables (t2m, d2m, sp) were missing, leading to errors.
Processing the new ERA5 format was a bottleneck.
The goal was to make extract.nc.ERA5 and met2CF.ERA5 functions flexible enough to support different ERA5 data formats, including both older and newer ensemble releases, as well as ERA5 reanalysis data. Generalizing this pipeline required a thorough understanding of ERA5 data structures and recent updates. To achieve this, I implemented robust format and timestep detection (hourly, 2-hourly, etc.) and checked for the presence of required variables before attempting to calculate specific humidity. If the necessary variables were missing, the calculation was safely skipped to prevent errors. For ensemble data with 4D, raster::brick fails or extracts only a slice unless explicitly adapted, so introduced logic to detect and correctly extract the ensemble dimension.
Added ERA5 Variable Mapping and Aligned Units to CF Standards ( PR #3604 )
Added an ERA5 column to the pecan_standard_met_table ( ERA5 is ECMWF's fifth-generation atmospheric reanalysis dataset providing global hourly meteorological data from 1940 to present). This enables PEcAn to correctly map ERA5 variable names (t2m, sp, d2m, strd, ssrd, tp, u10, v10, swvl1) to standardized meteorological variables.
Standardized units according to CF canonical standards in both the pecan_standard_met_table and standard_vars.csv, ensuring consistency across data sources.
AmeriFlux Meteorological Ensemble Processing with ERA5 Fallback ( PR # 3586 )
I developed the AmeriFlux_met_ensemble function for processing AmeriFlux meteorological data - - AmeriFlux is a network analogous to weather stations, but designed to track the exchange of CO2, H2O, and other gases between the atmosphere and ecosystems. The function encapsulates a comprehensive pipeline for processing AmeriFlux meteorological data, using ERA5 as a fallback for missing radiation and soil moisture data that is required to fill data gaps. The pipeline includes automated statistical modeling for gap-filling and ensemble generation.
Enhanced Meteorological Downscaling ( PR #3547 )
In this update, I worked on improving the meteorological (met) downscaling process. By understanding the dynamics and relations among variables.
The goal was twofold: expand the set of supported variables and resolve critical issues including a NetCDF writing bug that caused data loss through incorrect column detection, units inconsistencies (e.g, specific_humidity using non-CF-compliant g/g instead of kg/kg units), statistical distribution problems where normal sampling with clamping altered mean/standard deviation properties, and processing errors that returned all NA values for specific_humidity and incorrectly overwrote temperature bounds.
Downscaling now includes soil temperature, relative humidity (RH), soil moisture, and photosynthetic photon flux density (PPFD), with improved air_temperature_max and air_temperature_min processing using Gaussian ensemble methods with bounds checking. These additions make the workflow more comprehensive and it would be helpful to plot some of the resulting time series to visualize the downscaled values.
My Contributions to the PEcAn Project can be found here : PR List
Future Improvements
A planned improvement is to refactor the AmeriFlux meteorological ensemble workflow to make it fully modular within an existing PEcAn.atmosphere::met.process workflow. The goal is to reduce code duplication, align with PEcAn conventions, and improve maintainability. This update will integrate steps like metgapfill_with_fallback to automatically handle missing AmeriFlux radiation and soil moisture data using ERA5/ERA5-Land fallback. Additional enhancements include using reusable helper functions, adding unit tests for each component, and updating documentation to support the new modular workflow.
Current sampling in extract_soil_gssurgo uses frequency-based mukey aggregation rather than true area-weighted spatial integration, introducing systematic bias in soil property estimates. A critical architectural refactoring will migrate from direct WFS queries to the SoilDB package, leveraging mukey.wcs for native 30-meter gSSURGO resolution and eliminating overlapping buffer artifacts. This transition addresses fundamental spatial autocorrelation issues while implementing proper area-weighted aggregation through get_SDA_property methods, ensuring statistically robust soil ensemble generation that preserves both spatial heterogeneity and compositional constraints inherent to pedometric data structures.
The current ic_process enforces single-source initialization workflows, creating incomplete biogeochemical state vectors when data sources exhibit complementary coverage domains (e.g, SoilGrids providing soil carbon without vegetation pools). A planned architectural enhancement addresses this limitation through multi-source integration capabilities, enabling the combination of heterogeneous initial condition data source into cohesive poolinitcond files.
Acknowledgements
Embarking on the Google Summer of Code journey has been an extraordinary experience, and I am deeply grateful to all my mentors who supported and guided me along the way. It’s been a wonderful opportunity to work and learn alongside some highly motivating and talented mentors. The number of things I learned on the way was absolutely insane, and the feedback was really valuable.
A special note of gratitude goes to the lead mentor David LeBauer whose support has been invaluable at every stage of this project and that kept me energized and to Chris Black, who carefully reviewed and provided insightful feedback on my pull requests. I am equally grateful to the folks in the PEcAn community for being open and responsive to my questions.


Congratulations Akash on an outstanding summer of contributions! 🚀 We are happy to have you as a member of the community, and really appreciate your contributions to the workflows for estimating and propagating uncertainty from soils, weather, and initial conditions. You went above and beyond not only in developing workflows, but also in resurrecting and refactoring a lot of our code. It’s been a pleasure to mentor you through GSoC, and I’m excited to continue working with you moving forward!